The Unique Power of FACS-Based CRISPR Screens: Finding True Regulators Behind Complex Cellular Phenotypes
The Unique Power of FACS-Based CRISPR Screens: Finding True Regulators Behind Complex Cellular Phenotypes

DuneX CRISPR Team
•
Aug 5, 2025
Why Traditional CRISPR Screens Miss the Most Interesting Biology
High-throughput CRISPR screening has reshaped how drug developers and academic researchers interrogate biology, offering a systematic and scalable way to map genes to function. Yet as impactful as proliferation-based or drug-survival CRISPR screens have been, they capture only a narrow slice of the biological information encoded within a cell. Much of modern biomedical research—particularly immunology, oncology, metabolic regulation, and cell signaling—is defined not by survival outcomes but by subtle, quantitative, and multidimensional phenotypes. These are phenotypes that unfold gradually, vary across cell states, and often escape detection in bulk assays.
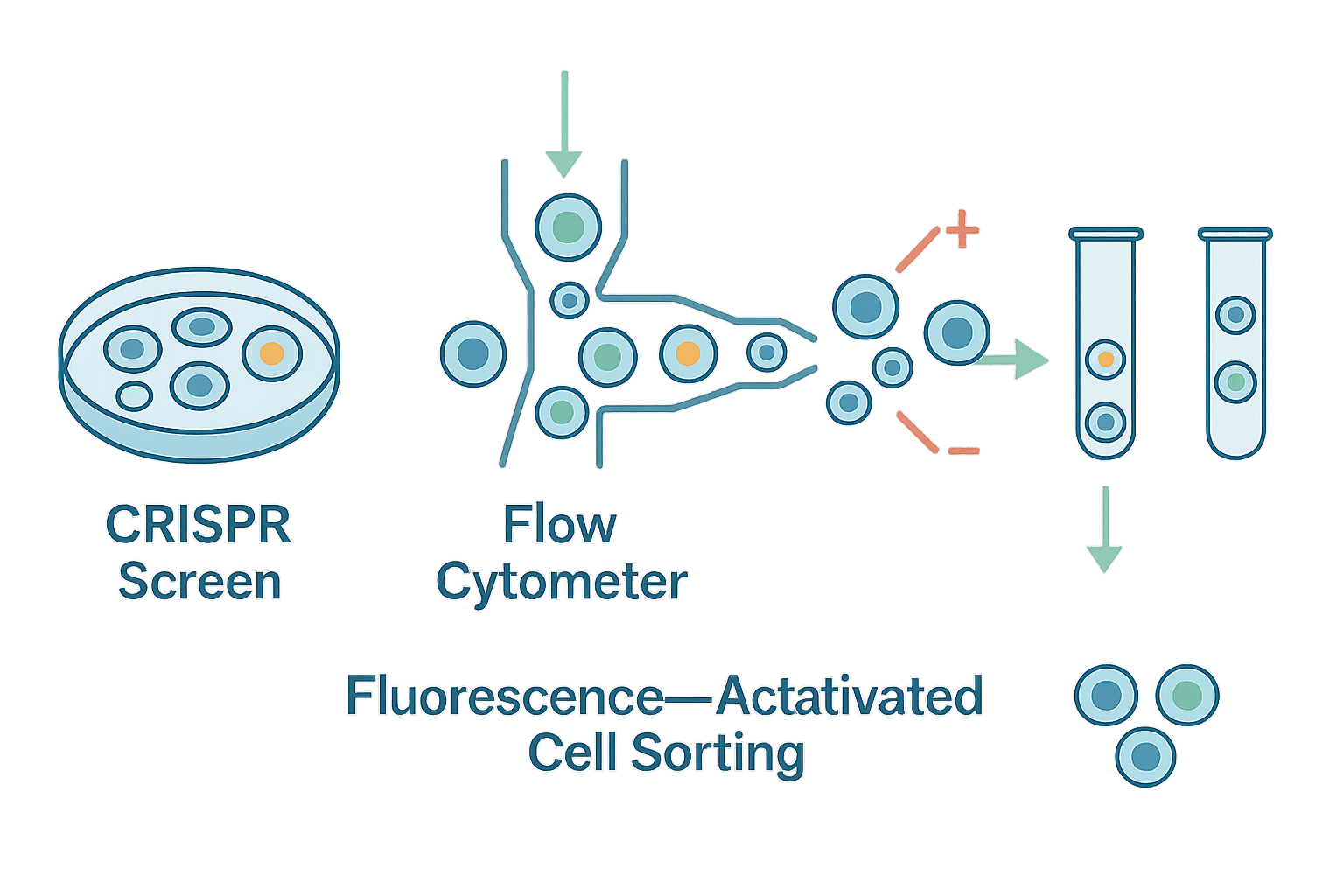
This is the space where FACS-based CRISPR screens distinguish themselves as uniquely powerful. By coupling pooled CRISPR perturbations with the resolution of flow cytometry, researchers can track fluorescence, protein abundance, signaling states, or intracellular markers at single-cell resolution, and then isolate specific subpopulations for sequencing. This approach provides a far richer and more nuanced readout than survival-based screens and has led to discoveries that would otherwise remain invisible.
CRISPR screens were originally popularized for viability studies after seminal demonstrations in human cells that established the method’s scalability and power (Shalem et al., 2014; Wang et al., 2014). These early screens produced sharp genetic-dependency maps and were instrumental in the rise of synthetic lethality–based drug development. But as CRISPR technology matured, it became clear that a substantial fraction of biologically meaningful signals—immune activation thresholds, receptor-expression shifts, transcription-factor dynamics, chromatin accessibility changes, and metabolic rewiring—cannot be captured through survival metrics alone. The transition to fluorescence-based selection represented a conceptual expansion: instead of asking “Which genes make cells live or die?” scientists could begin asking, “Which genes shape the architecture of this phenotype?”
How Flow Cytometry Unlocks Single-Cell Resolution in CRISPR Screens
Flow cytometry has long been valued for its ability to capture single-cell heterogeneity, particularly in immunology. When integrated with pooled CRISPR libraries, this capability becomes transformative. Rather than working with population averages, researchers can gate cells based on subtle fluorescence differences—sometimes a two-fold change, sometimes even less—and isolate the top or bottom percentile of cells exhibiting a phenotype of interest. Sequencing then reveals which gene perturbations drive those shifts. This is fundamentally different from bulk methods, where nuanced signaling differences are masked by noise.
A central strength of FACS-based CRISPR screens lies in their ability to work with continuous phenotypes, not just binary ones. Cellular biology rarely presents all-or-nothing behaviors; more often, phenotypes move along gradients. Receptor upregulation, pathway activation, mitochondrial potential, calcium flux, cytokine secretion, or transcription-factor nuclear localization each occupy a spectrum. A drug may increase PD-L1 expression by 40%, reduce NF-κB reporter activity by 25%, or shift a metabolic ratio across a broad range. In bulk assays, these changes dilute out. With FACS gating, researchers can select specific fluorescence-defined subpopulations and read out the gene perturbations that reshape the distribution.
Capturing Multi-Layered Gene Networks Rather Than Single-Gene Effects
This capacity becomes even more valuable when studying multi-gene regulatory networks, which rarely collapse into simple linear pathways. Consider T cell activation, an area that has benefited significantly from FACS-based CRISPR screening. Signal transduction through the TCR engages dozens of transcriptional regulators, chromatin remodelers, metabolic checkpoints, and protein-turnover mechanisms. Shifrut et al. (2018) demonstrated that genome-wide CRISPR screens in primary human T cells can reveal regulators of activation, cytokine production, and functional exhaustion—insights that rely on fluorescence-based gating of markers like CD69, CD25, or IFN-γ. Without FACS selection, many regulators would appear indistinguishable from background noise. These findings now guide CAR-T engineering strategies, highlighting genes that modulate activation thresholds or exhaustion resistance.
Reporter-Based CRISPR Screens: Measuring Pathways That Survival Screens Cannot See
Another decisive advantage of FACS-based screens is their compatibility with reporter constructs, which are now widely used in functional genomics and drug discovery. Signaling reporters—such as NFAT-GFP, NF-κB-luciferase, STAT-pathway sensors, or synthetic Notch-based logic gates—allow researchers to quantify pathway engagement dynamically. By enriching for cells in the highest or lowest fluorescence bins, screens can reveal upstream regulators of pathway amplitude or duration. Similar approaches apply to reporters of DNA-damage signaling, ER stress, mitochondrial activity, antigen presentation, metabolic flux, and more. These reporters convert otherwise intangible intracellular changes into measurable fluorescence, which FACS can cleanly resolve.
The importance of reporter-based screening is highlighted in the study of drug mechanisms. While viability-based screens reveal essential genes or sensitizers, they often fail to resolve early and subtle pharmacodynamic responses. In contrast, FACS-based CRISPR screens can identify regulatory bottlenecks at intermediate nodes of a drug’s mechanism. For instance, PARP inhibitor studies have uncovered regulators of DNA-repair pathway choice, including shieldin and POLQ, which modulate the cellular response to replication stress (Zimmermann et al., 2018). In more complex phenotypes, such as immune checkpoint regulation, FACS screens have successfully identified transcriptional and chromatin regulators that subtly modulate PD-L1 levels, providing insights into tumor immune evasion.
Avoiding Survival Bias and Revealing True Biological Modulators
A related strength of FACS-based CRISPR screens is that they operate outside the constraints of cell survival bias, which often distorts results in proliferation-based screens. Many biologically important perturbations do not kill cells; they simply alter state, trajectory, or responsiveness. Survival screens intrinsically filter out slower-growing cells and amplify fast-growing ones, independently of the phenotype being studied. FACS avoids this bias entirely, letting researchers enrich for cells based purely on the phenotype of interest.
From a practical standpoint, the combination of CRISPR and FACS imposes technical challenges—particularly in preserving library representation while sorting sufficient numbers of cells. Maintaining coverage across bins is essential, and sorting too stringently can collapse representation. But these concerns are readily managed with proper experimental design, including pre-sorting timepoint sequencing, moderate bin sizes (e.g., top and bottom 5–10%), and careful optimization of reporter expression. Groups performing high-quality CRISPR screens often treat coverage as the single most important determinant of success, a principle equally true for FACS-based approaches (Doench, 2018).
Interpreting Data: Continuous Phenotypes and Stronger Biological Signals
As with all CRISPR modalities, data interpretation relies on models that integrate guide-level and gene-level effects. MAGeCK, CasTLE, and other statistical frameworks remain foundational (Meyers et al., 2017), and they adapt well to FACS-based count distributions. What distinguishes fluorescence-based screens is the shape of the signal: hits often show strong, coherent enrichment across multiple sgRNAs, and pathway-level relationships emerge more clearly than in survival-focused studies.
Where FACS-Based Screens Are Heading Next
Looking ahead, FACS-based CRISPR screening is increasingly complemented by Perturb-seq, which layers single-cell transcriptomic profiling on top of pooled gene perturbations. While FACS identifies the regulatory genes, Perturb-seq provides mechanistic detail about downstream transcriptional consequences. The two workflows form a natural pair: FACS identifies the modulators; Perturb-seq explains how they work. Emerging technologies, including high-parameter spectral cytometry, intracellular phospho-specific staining protocols, and multiomic flow platforms, promise to expand the resolution further. These advances may ultimately allow CRISPR screens to track dozens of intracellular states simultaneously.
Despite these technological expansions, the foundational appeal of FACS-based CRISPR screening remains elegantly simple: it allows researchers to focus on what actually changes in cells, not only on whether the cells live or die. Biology is not a binary system, and the most therapeutically relevant phenotypes frequently occupy the gradients in between. By enabling high-throughput genetic interrogation of these subtle, graded phenotypes, fluorescence-based CRISPR screens provide drug developers with a profoundly powerful way to reveal true regulators—those elusive hits that govern cellular behavior in complex, multilayered systems.
References
Shalem, O. et al. (2014) Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87.
Wang, T. et al. (2014) Genetic screens in human cells using the CRISPR–Cas9 system. Science 343, 80–84.
Horlbeck, M. A. et al. (2016) Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. Nat. Methods 13, 971–974.
Zimmermann, M. et al. (2018) CRISPR screens identify genomic factors that determine PARP inhibitor sensitivity. Cell 173, 1–15.
Shifrut, E. et al. (2018) Genome-wide CRISPR screens in primary human T cells reveal key regulators of immune function. Cell 175, 1958–1971.
Konermann, S. et al. (2015) Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588.
Meyers, R. M. et al. (2017) Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens. Nat. Genet. 49, 1779–1784.
Doench, J. G. (2018) Am I ready for CRISPR? A user’s guide to genetic screens. Nat. Rev. Genet. 19, 67–80.
Why Traditional CRISPR Screens Miss the Most Interesting Biology
High-throughput CRISPR screening has reshaped how drug developers and academic researchers interrogate biology, offering a systematic and scalable way to map genes to function. Yet as impactful as proliferation-based or drug-survival CRISPR screens have been, they capture only a narrow slice of the biological information encoded within a cell. Much of modern biomedical research—particularly immunology, oncology, metabolic regulation, and cell signaling—is defined not by survival outcomes but by subtle, quantitative, and multidimensional phenotypes. These are phenotypes that unfold gradually, vary across cell states, and often escape detection in bulk assays.
This is the space where FACS-based CRISPR screens distinguish themselves as uniquely powerful. By coupling pooled CRISPR perturbations with the resolution of flow cytometry, researchers can track fluorescence, protein abundance, signaling states, or intracellular markers at single-cell resolution, and then isolate specific subpopulations for sequencing. This approach provides a far richer and more nuanced readout than survival-based screens and has led to discoveries that would otherwise remain invisible.
CRISPR screens were originally popularized for viability studies after seminal demonstrations in human cells that established the method’s scalability and power (Shalem et al., 2014; Wang et al., 2014). These early screens produced sharp genetic-dependency maps and were instrumental in the rise of synthetic lethality–based drug development. But as CRISPR technology matured, it became clear that a substantial fraction of biologically meaningful signals—immune activation thresholds, receptor-expression shifts, transcription-factor dynamics, chromatin accessibility changes, and metabolic rewiring—cannot be captured through survival metrics alone. The transition to fluorescence-based selection represented a conceptual expansion: instead of asking “Which genes make cells live or die?” scientists could begin asking, “Which genes shape the architecture of this phenotype?”
How Flow Cytometry Unlocks Single-Cell Resolution in CRISPR Screens
Flow cytometry has long been valued for its ability to capture single-cell heterogeneity, particularly in immunology. When integrated with pooled CRISPR libraries, this capability becomes transformative. Rather than working with population averages, researchers can gate cells based on subtle fluorescence differences—sometimes a two-fold change, sometimes even less—and isolate the top or bottom percentile of cells exhibiting a phenotype of interest. Sequencing then reveals which gene perturbations drive those shifts. This is fundamentally different from bulk methods, where nuanced signaling differences are masked by noise.
A central strength of FACS-based CRISPR screens lies in their ability to work with continuous phenotypes, not just binary ones. Cellular biology rarely presents all-or-nothing behaviors; more often, phenotypes move along gradients. Receptor upregulation, pathway activation, mitochondrial potential, calcium flux, cytokine secretion, or transcription-factor nuclear localization each occupy a spectrum. A drug may increase PD-L1 expression by 40%, reduce NF-κB reporter activity by 25%, or shift a metabolic ratio across a broad range. In bulk assays, these changes dilute out. With FACS gating, researchers can select specific fluorescence-defined subpopulations and read out the gene perturbations that reshape the distribution.

Capturing Multi-Layered Gene Networks Rather Than Single-Gene Effects
This capacity becomes even more valuable when studying multi-gene regulatory networks, which rarely collapse into simple linear pathways. Consider T cell activation, an area that has benefited significantly from FACS-based CRISPR screening. Signal transduction through the TCR engages dozens of transcriptional regulators, chromatin remodelers, metabolic checkpoints, and protein-turnover mechanisms. Shifrut et al. (2018) demonstrated that genome-wide CRISPR screens in primary human T cells can reveal regulators of activation, cytokine production, and functional exhaustion—insights that rely on fluorescence-based gating of markers like CD69, CD25, or IFN-γ. Without FACS selection, many regulators would appear indistinguishable from background noise. These findings now guide CAR-T engineering strategies, highlighting genes that modulate activation thresholds or exhaustion resistance.
Reporter-Based CRISPR Screens: Measuring Pathways That Survival Screens Cannot See
Another decisive advantage of FACS-based screens is their compatibility with reporter constructs, which are now widely used in functional genomics and drug discovery. Signaling reporters—such as NFAT-GFP, NF-κB-luciferase, STAT-pathway sensors, or synthetic Notch-based logic gates—allow researchers to quantify pathway engagement dynamically. By enriching for cells in the highest or lowest fluorescence bins, screens can reveal upstream regulators of pathway amplitude or duration. Similar approaches apply to reporters of DNA-damage signaling, ER stress, mitochondrial activity, antigen presentation, metabolic flux, and more. These reporters convert otherwise intangible intracellular changes into measurable fluorescence, which FACS can cleanly resolve.
The importance of reporter-based screening is highlighted in the study of drug mechanisms. While viability-based screens reveal essential genes or sensitizers, they often fail to resolve early and subtle pharmacodynamic responses. In contrast, FACS-based CRISPR screens can identify regulatory bottlenecks at intermediate nodes of a drug’s mechanism. For instance, PARP inhibitor studies have uncovered regulators of DNA-repair pathway choice, including shieldin and POLQ, which modulate the cellular response to replication stress (Zimmermann et al., 2018). In more complex phenotypes, such as immune checkpoint regulation, FACS screens have successfully identified transcriptional and chromatin regulators that subtly modulate PD-L1 levels, providing insights into tumor immune evasion.
Avoiding Survival Bias and Revealing True Biological Modulators
A related strength of FACS-based CRISPR screens is that they operate outside the constraints of cell survival bias, which often distorts results in proliferation-based screens. Many biologically important perturbations do not kill cells; they simply alter state, trajectory, or responsiveness. Survival screens intrinsically filter out slower-growing cells and amplify fast-growing ones, independently of the phenotype being studied. FACS avoids this bias entirely, letting researchers enrich for cells based purely on the phenotype of interest.
From a practical standpoint, the combination of CRISPR and FACS imposes technical challenges—particularly in preserving library representation while sorting sufficient numbers of cells. Maintaining coverage across bins is essential, and sorting too stringently can collapse representation. But these concerns are readily managed with proper experimental design, including pre-sorting timepoint sequencing, moderate bin sizes (e.g., top and bottom 5–10%), and careful optimization of reporter expression. Groups performing high-quality CRISPR screens often treat coverage as the single most important determinant of success, a principle equally true for FACS-based approaches (Doench, 2018).
Interpreting Data: Continuous Phenotypes and Stronger Biological Signals
As with all CRISPR modalities, data interpretation relies on models that integrate guide-level and gene-level effects. MAGeCK, CasTLE, and other statistical frameworks remain foundational (Meyers et al., 2017), and they adapt well to FACS-based count distributions. What distinguishes fluorescence-based screens is the shape of the signal: hits often show strong, coherent enrichment across multiple sgRNAs, and pathway-level relationships emerge more clearly than in survival-focused studies.
Where FACS-Based Screens Are Heading Next
Looking ahead, FACS-based CRISPR screening is increasingly complemented by Perturb-seq, which layers single-cell transcriptomic profiling on top of pooled gene perturbations. While FACS identifies the regulatory genes, Perturb-seq provides mechanistic detail about downstream transcriptional consequences. The two workflows form a natural pair: FACS identifies the modulators; Perturb-seq explains how they work. Emerging technologies, including high-parameter spectral cytometry, intracellular phospho-specific staining protocols, and multiomic flow platforms, promise to expand the resolution further. These advances may ultimately allow CRISPR screens to track dozens of intracellular states simultaneously.
Despite these technological expansions, the foundational appeal of FACS-based CRISPR screening remains elegantly simple: it allows researchers to focus on what actually changes in cells, not only on whether the cells live or die. Biology is not a binary system, and the most therapeutically relevant phenotypes frequently occupy the gradients in between. By enabling high-throughput genetic interrogation of these subtle, graded phenotypes, fluorescence-based CRISPR screens provide drug developers with a profoundly powerful way to reveal true regulators—those elusive hits that govern cellular behavior in complex, multilayered systems.
References
Shalem, O. et al. (2014) Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87.
Wang, T. et al. (2014) Genetic screens in human cells using the CRISPR–Cas9 system. Science 343, 80–84.
Horlbeck, M. A. et al. (2016) Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. Nat. Methods 13, 971–974.
Zimmermann, M. et al. (2018) CRISPR screens identify genomic factors that determine PARP inhibitor sensitivity. Cell 173, 1–15.
Shifrut, E. et al. (2018) Genome-wide CRISPR screens in primary human T cells reveal key regulators of immune function. Cell 175, 1958–1971.
Konermann, S. et al. (2015) Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588.
Meyers, R. M. et al. (2017) Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens. Nat. Genet. 49, 1779–1784.
Doench, J. G. (2018) Am I ready for CRISPR? A user’s guide to genetic screens. Nat. Rev. Genet. 19, 67–80.

Copyright © 2025 DuneX Biosciences. All rights reserved. | +1-(415).463.0365 | info@dunexbio.com | 25801 Industrial Blvd Suite 100, Hayward, CA 94545

Copyright © 2025 DuneX Biosciences. All rights reserved. | +1-(415).463.0365 | info@dunexbio.com | 25801 Industrial Blvd Suite 100, Hayward, CA 94545

Copyright © 2025 DuneX Biosciences.
All rights reserved.
+1-(415).463.0365 | info@dunexbio.com |
25801 Industrial Blvd Suite 100, Hayward, CA 94545

Copyright © 2025 DuneX Biosciences. All rights reserved. | +1-(415).463.0365 | info@dunexbio.com |
25801 Industrial Blvd Suite 100, Hayward, CA 94545
