Key outcome
The best-performing clone reached an apparent yeast-display Kd of 17.12 nM in the EGF titration panel. Additional variants created a useful ranking set rather than a single isolated hit.

Case Study - Rapid Binder Discovery Against EGF
6 sequence-confirmed binders identified from a compact 53-aa target, with the best clone showing an apparent yeast-display Kd of 17.12 nM. This pilot-scale yeast display case study demonstrates FACS enrichment, clone recovery, and dose-dependent binding against a compact, high-background antigen.
Results
Using the Nova-Intermediate-Coop library, the demo produced a measurable binder panel despite elevated secondary/background signal, supporting both the discovery workflow and next-round optimization strategy.
The best-performing clone reached an apparent yeast-display Kd of 17.12 nM in the EGF titration panel. Additional variants created a useful ranking set rather than a single isolated hit.
This case study demonstrates a complete early discovery loop: antigen staining, FACS enrichment, clone recovery, and clone-level titration. For client projects, the same structure can be expanded with alternate labels, competition sorting, off-rate selection, soluble reformatting, and orthogonal binding assays.
For small antigens like EGF, the most important next optimization is often reagent-background control, including alternate fluorophore/streptavidin formats and negative depletion against secondary reagents.
Open Dataset
Review the original Yeast Display Demo repository for raw FCS files, gating strategy, NGS material, and clone binding curve assets behind this case study.
Program Goal
EGF is a compact 53-amino-acid protein, making it a useful demonstration target for workflows where antigen size, labeling format, and secondary-reagent background can all affect apparent enrichment.
The objective was not only to show enrichment, but to generate a small panel of individual binders that could be sequenced, re-tested, and ranked by concentration titration.
EGF-like targets can be sensitive to label orientation and streptavidin-PE background. The workflow emphasized gated populations, round-by-round enrichment, and secondary-background awareness.
The resulting binders provide a practical launch point for confirmatory assays, alternate labeling formats, off-rate selection, or affinity maturation when tighter binding is required.
Approach
The workflow combines yeast display selection with FACS-based gates and a final titration panel, converting an enriched population into individual clone-level binding data.
The synthetic scFv-format library was screened to identify EGF-reactive variants while maintaining a practical path to downstream sequence analysis and reformatting.
Yeast cells were gated for clean populations and sorted across R1-R4. Enrichment gates prioritized antigen-positive events while tracking background.
Recovered clones were tested across 1-300 nM EGF to generate dose-response curves and estimate apparent yeast-display binding strength.
Selection Panels
Round-level FACS plots show how antigen-positive populations were tracked and sorted during the discovery workflow.
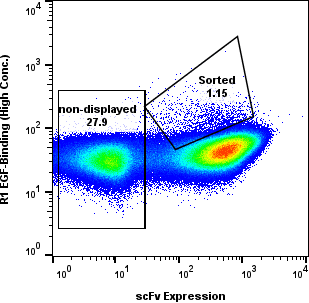
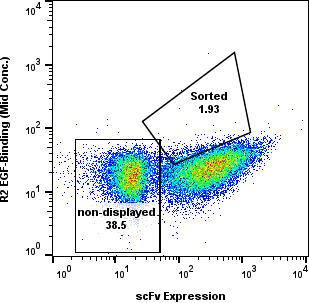
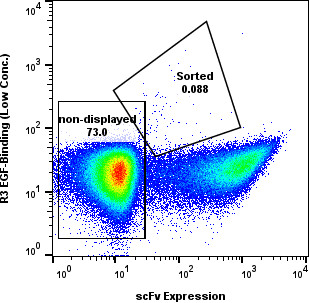
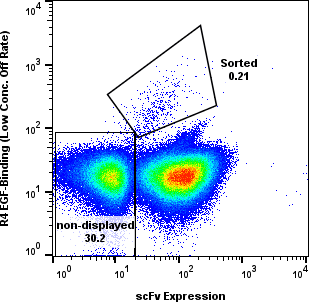
Dose Series
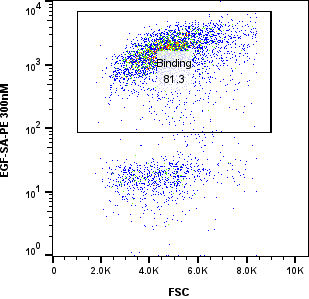
The tabs below let technical reviewers inspect the raw Variant 1 FACS gate behavior from 300 nM down to 1 nM.
Typical Timeline
Actual timing depends on antigen availability, label format, confirmation depth, and whether soluble reformatting is included.
Review antigen format, labeling strategy, control reagents, and expected downstream use.
Run sequential enrichment rounds with population gates, background checks, and sorted libraries.
Pick, sequence, and re-test enriched clones to identify unique candidate binders.
Generate titration curves, apparent Kd estimates, and a prioritized binder panel.
Share your target format, project goal, and desired binder profile. We’ll help map the fastest practical path toward a focused yeast display discovery campaign.
Contact Us